Integrate single-cell RNA-Seq datasets in R using Seurat (CCA) | Detailed Seurat Workflow Tutorial - YouTube
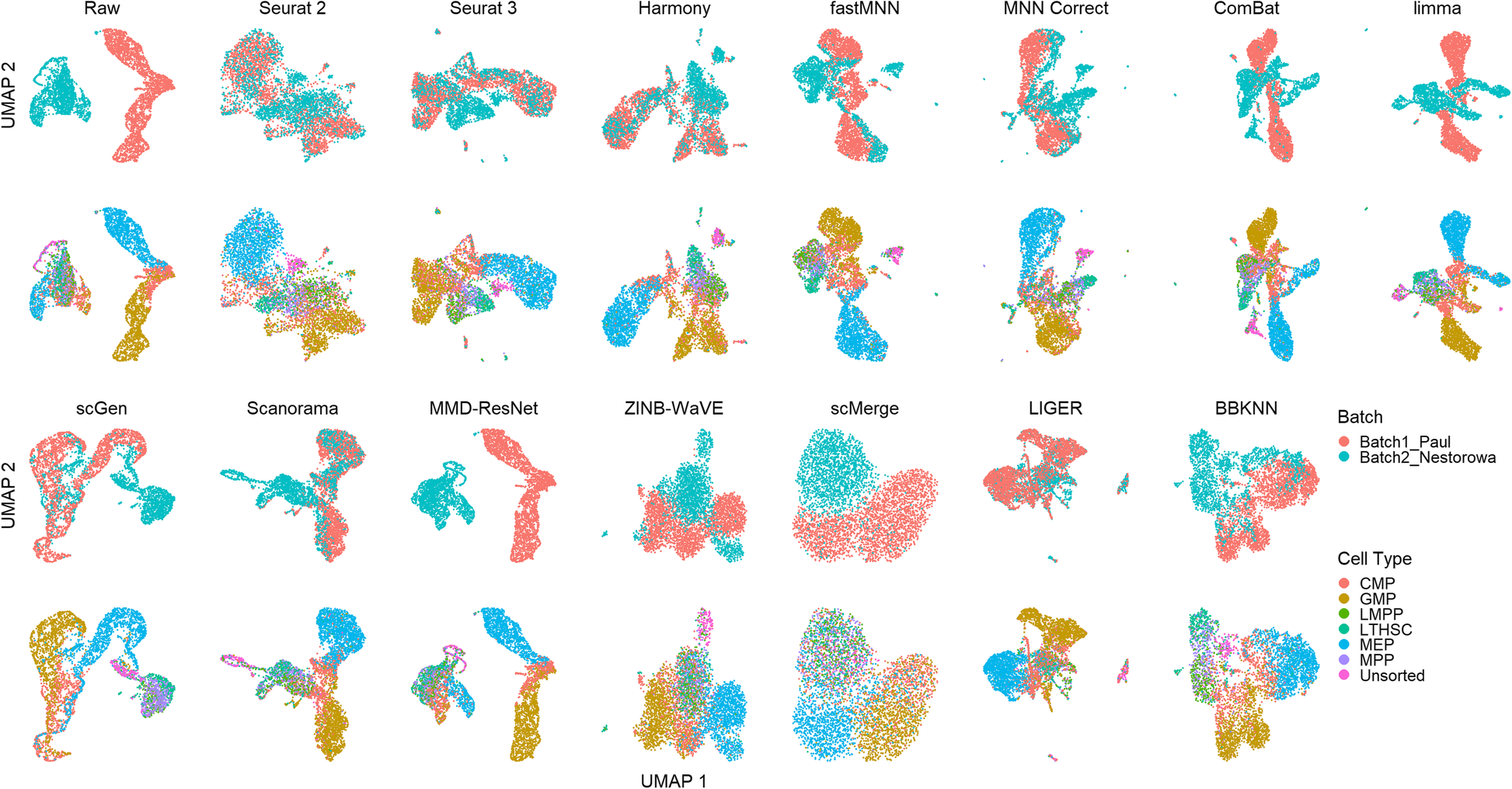
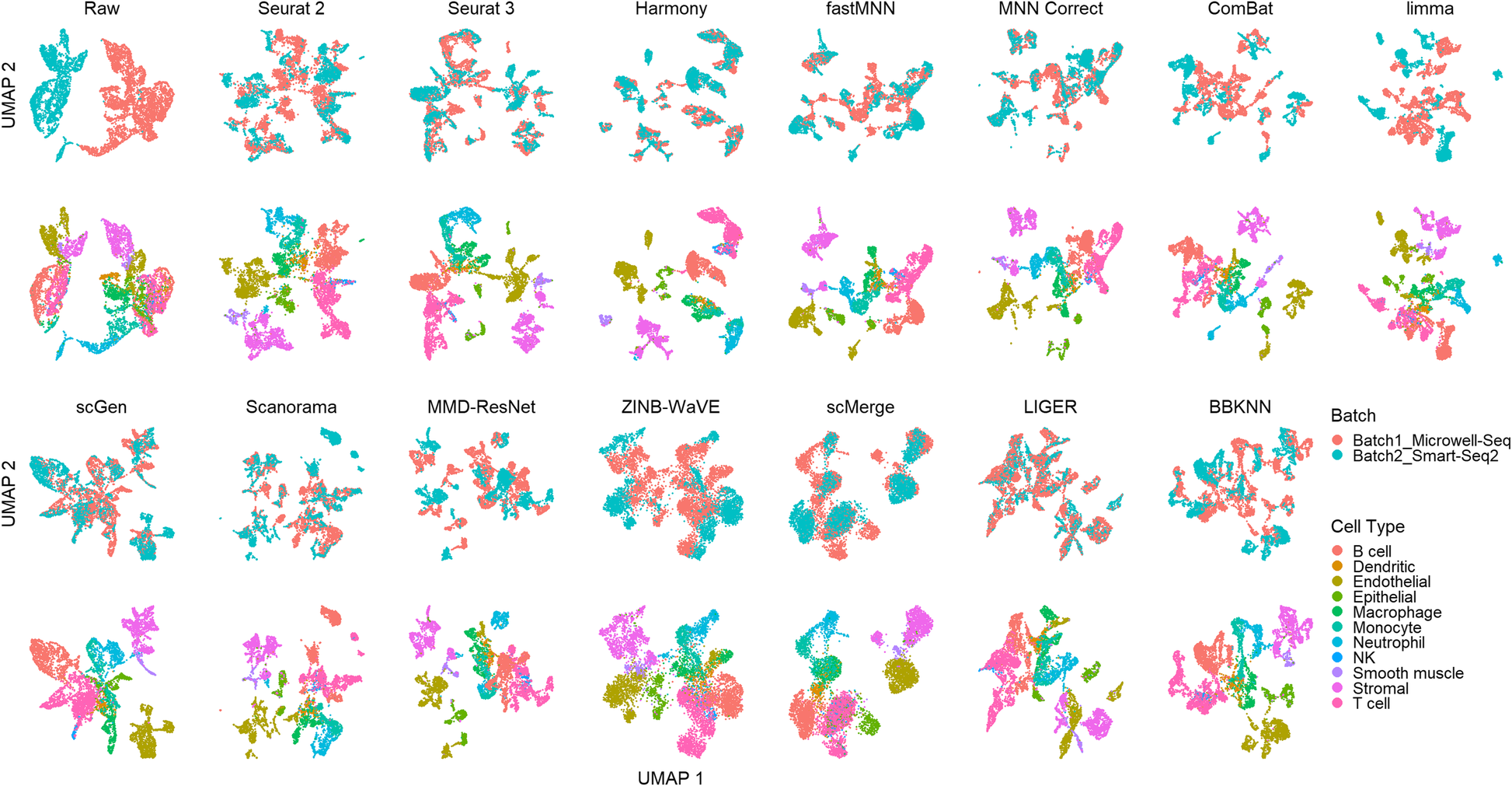
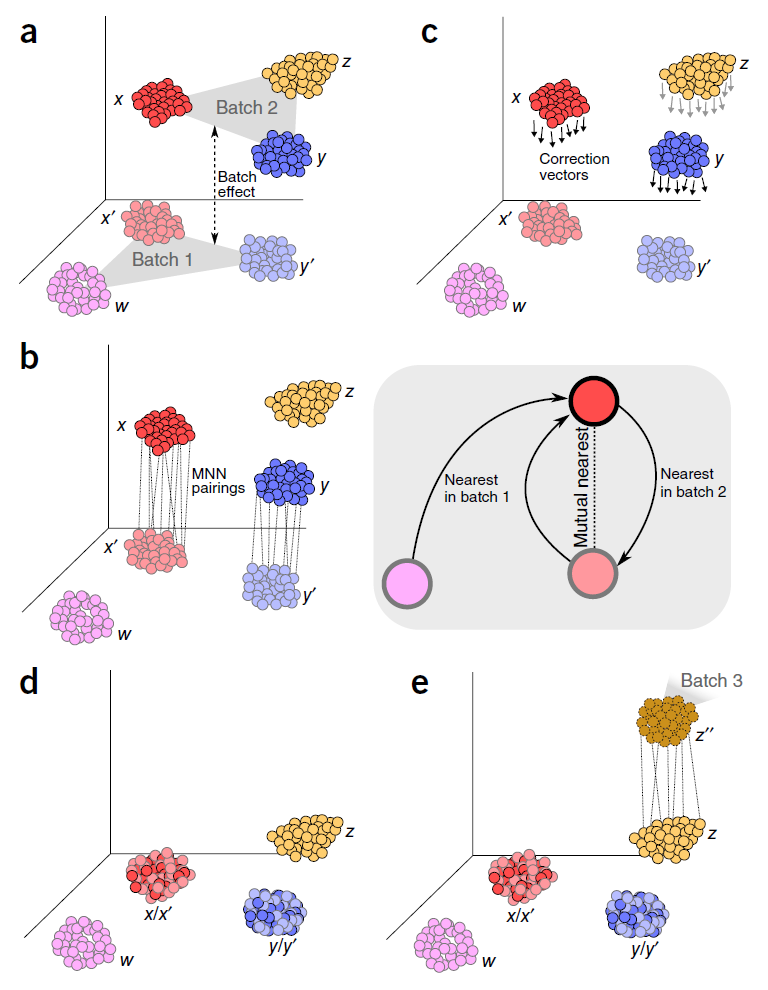
A benchmark of batch-effect correction methods for single-cell RNA sequencing data | Genome Biology | Full Text

Comparisons of data integration methods for batch effect correction for... | Download Scientific Diagram
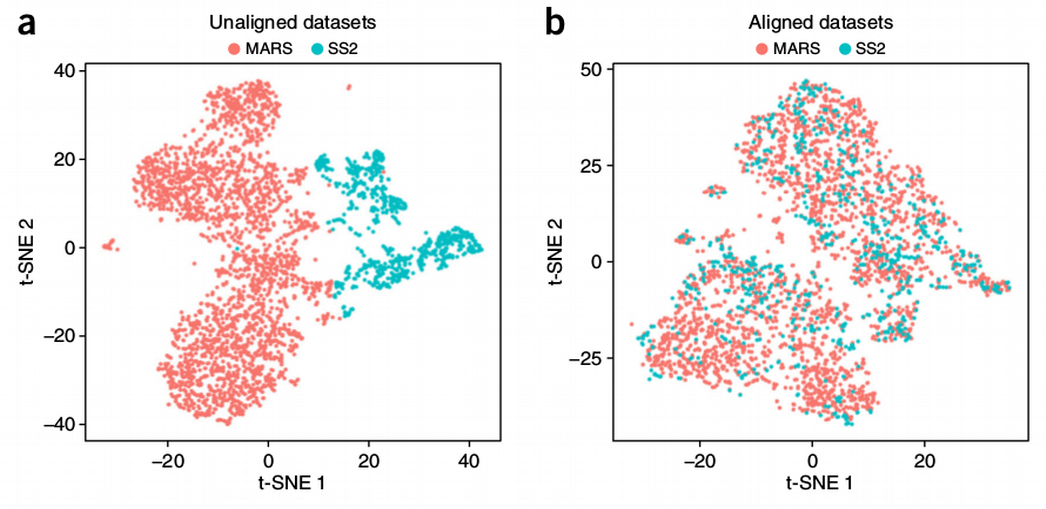
How to Batch Correct Single Cell. Comparing batch correction methods for… | by Nikolay Oskolkov | Towards Data Science

A benchmark of batch-effect correction methods for single-cell RNA sequencing data | Genome Biology | Full Text

Comparison of Scanpy-based algorithms to remove the batch effect from single-cell RNA-seq data | Cell Regeneration | Full Text

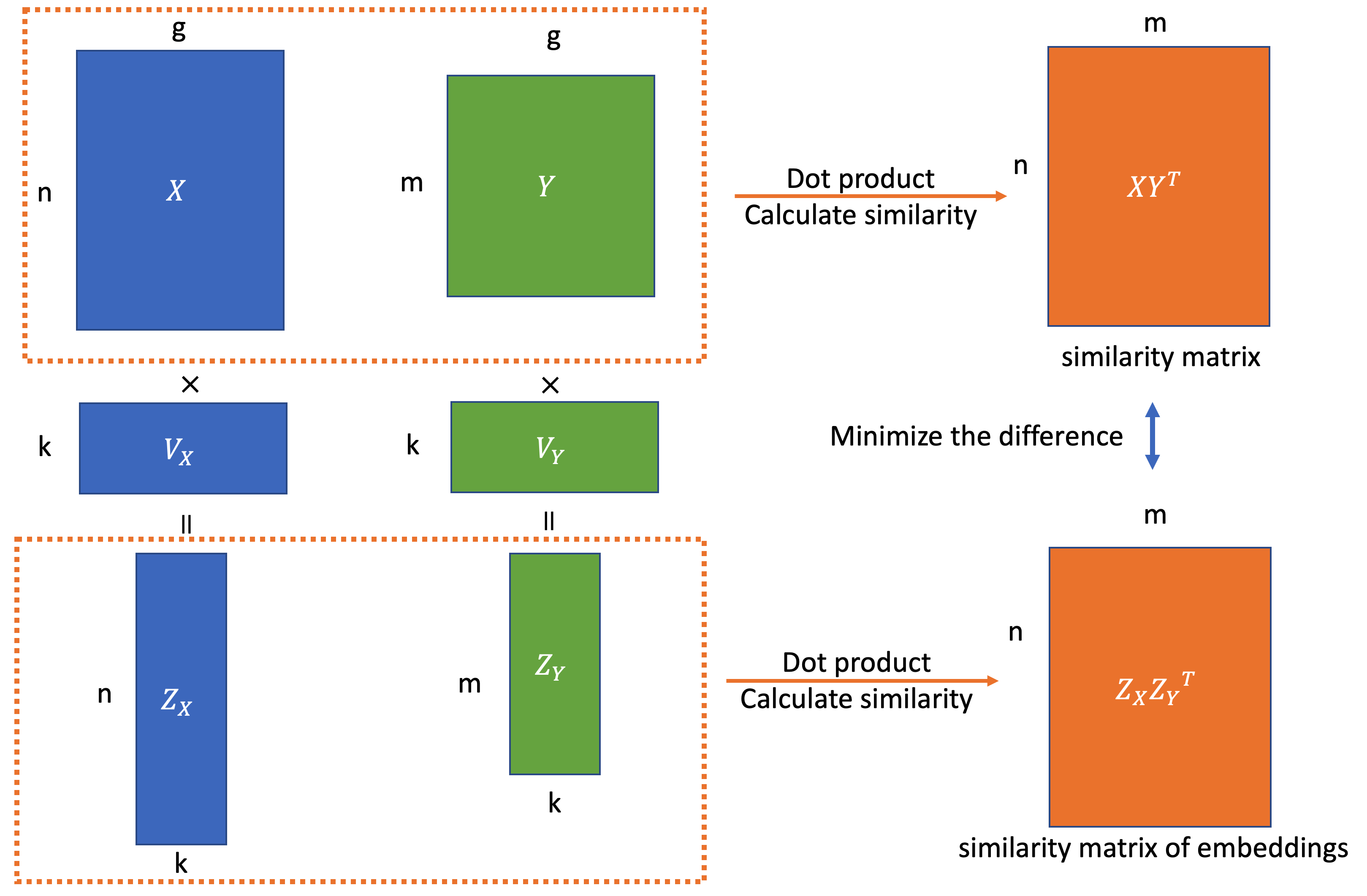
Batch alignment of single-cell transcriptomics data using deep metric learning | Nature Communications

A benchmark of batch-effect correction methods for single-cell RNA sequencing data. - Abstract - Europe PMC
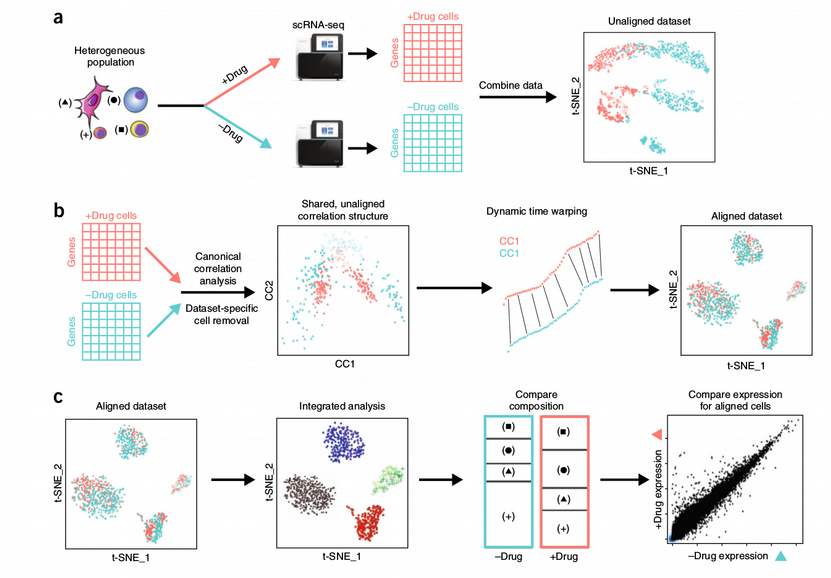
How to Batch Correct Single Cell. Comparing batch correction methods for… | by Nikolay Oskolkov | Towards Data Science